This blog post is the first of three to discuss the Genome Archive (GenArk) assembly hubs. This first post discusses accessing the GenArk hubs, the second post gives examples of using the data, and the third post describes the technical infrastructure behind the hubs.
Let’s start with a real-world story: imagine you are a researcher working on zebrafish, but you are using an alternative strain with unique polymorphic properties. You have a desire to do CRISPR on your particular zebrafish and you already have a FASTA file for the genome assembled into chromosomes, but have no annotations or way to visualize the data yet.
One option to visualize your FASTA would be to independently create a UCSC Assembly Track Hub to work on your zebrafish. Or now that UCSC has developed the Genome Archive (GenArk) system, when you submit your assembly into NCBI’s assembly database, you could contact us directly and request we generate the browser for you behind the scenes. This happened for a specific lab, where they submitted their specific TD5 zebrafish assembly to NCBI, https://www.ncbi.nlm.nih.gov/assembly/GCA_018400075.1, and the result after contacting us was a new assembly hub that could be easily loaded at UCSC with the following link: https://genome.ucsc.edu/h/GCA_018400075.1 In this case, the team at UCSC even helped generate liftOver alignment files between the UCSC zebrafish in this new TD5 zebrafish GenArk Public Hub addition, aiding identification of lifting annotations to the new browser.
So what are the GenArk Assembly hubs?
GenArk hubs are a collection of data files externally hosted from the main UCSC data website enabling browsing new genomes. GenArk genomes have NCBI Genbank assembly accessions starting with either GCA or GCF and the browsers allow visualizing and attaching laboratory-generated data. New software also enables UCSC to dynamically turn on query servers to search GenArk hubs with DNA sequences or test PCR primer pairs. GenArk hubs are part of the UCSC Public Hubs list where UCSC can update the data files with pipelines.
ACCESSING THE DATA
How do I access GenArk Assembly Hubs?
There are multiple ways to access the GenArk hubs, including searching the UCSC Genome Browser’s main gateway page, building direct links to NCBI GCA/GCF assembly accessions, searching the UCSC Public Hubs page, and navigating directly to individual taxonomic group pages.
Browser Gateway Page
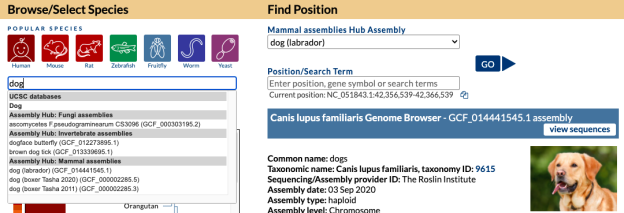
The easiest way to find GenArk hubs is to search the species name on the Browser Gateway.
On the Gateway page in the top left box you can search a term such as “dog” and find all the genomes both hosted in our internal databases and in external Public Hubs that have dog in the name. In this image, a search for “dog” returns a top “Dog” match (UCSC database) as well as results for several species in Assembly Track Hubs that match on the term “dog” with the specific labrador dog breed selected from the GenArk Mammal Assemblies Hub (GCF_014441545.1).
Direct GCA/GCF Accession Links
In the situation where you may know the GCF/GCA identifier for an assembly, you can also search that term on the Gateway page or build a short link to directly load the hub. Links to UCSC with a hub (“/h/”) address, such as https://genome.ucsc.edu/h/GCF_000698965.1 will attempt to find and attach a matching final GCF-value, which originates from the NCBI accession, in this case, for an African ostrich assembly. If you don’t find a match, read more below about contacting us.
Public Hubs Page
Another place to find GenArk hubs is on the Public Hubs page where you can enter various terms, like “ostrich”: https://genome.ucsc.edu/cgi-bin/hgHubConnect?hubSearchTerms=ostrich, You can expand the “Search details” to examine matching results. To load a desired hub, use a right-click to display an “Open this assembly” pop-up, or an option to configure individual track settings.
Genomes Menu
Another option to gain an overview of all the GenArk hubs is to click the “Genome Archive GenArk” link available under the “Genomes” menu.
This Genomes menu link will open the GenArk homepage. On the GenArk homepage, a variety of links exist including the line,“Please note: text file listing of 1,600 NCBI/VGP genome assembly hubs.” Clicking that link will open a single text file that lists all available hubs allowing a quick overview: https://hgdownload.soe.ucsc.edu/hubs/UCSC_GI.assemblyHubList.txt
Individual Taxonomic Pages
The GenArk homepage also has links to specific taxonomic groupings hub pages, such as mammals, fishes, or fungi. For instance, a “birds” link, https://hgdownload.soe.ucsc.edu/hubs/birds/index.html, brings you to a webpage with links to launch browsers, along with links to other details for each assembly.
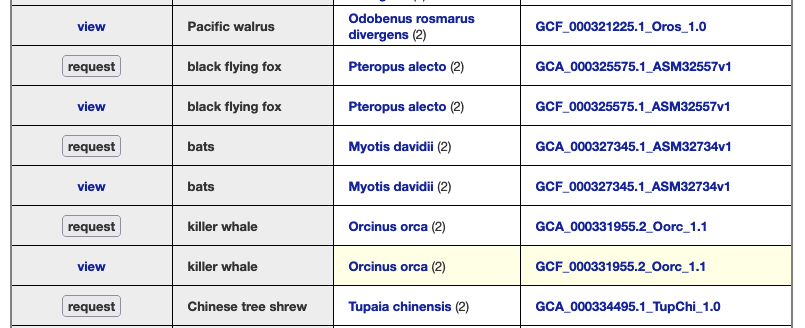
These taxonomic group pages, such as this image of the bird’s page, have links to launch the browser (2nd column: common name and view in browser) and links to the source files (4th column: NCBI assembly).
Access to these taxonomic group pages is also available from the Public Hubs page.
By going to the Description column on the Public Hubs page you can click a link (Bird genome assemblies) to end up at the related taxonomic grouping page. Also of note that on the Public Hubs page, you can click a [+] plus button to expand the list of Assemblies and click one of the GCA/GCF accession links to directly load an assembly.
What if I don’t find my Assembly in the GenArk collection?
If the assembly of interest is not found, please visit our assembly request page. Search that page for your assembly, if there is a “view” link you can launch the existing genome browser. Otherwise, click the “request” button to fill out a form to add your genome of interest.
The assembly request page does require there to already be an existing GCA/GCF identifier. You can also always email us at our public mailing-list genome@ucsc.soe.edu to request we add the assembly to the GenArk collection. This archived mailing-list is searchable from links on our contacts page, http://genome.ucsc.edu/contacts.html. Alternatively, if you don’t want your request to be public, you can email our private internal mailing-list at genome-www@soe.ucsc.edu.
What if my assembly doesn’t have a GCA/GCF NCBI accession?
If NCBI does not have a GCA/GCF accession for your assembly then our scripts will not be able to pull the data and generate the GenArk hub. You will need to deposit the assembly at NCBI and notify us once the assembly has become available. You can find directions at NCBI for how to submit new genomes: https://www.ncbi.nlm.nih.gov/assembly/docs/submission/
The next blog post in this series will provide examples of using the GenArk hubs, such as the BLAT and PCR tools that are available, or how you can send DNA of any Assembly Hubs to External Tools for processing. The final post examines the infrastructure behind the hubs.
This entry written by Brian Lee. If after reading this blog post you have any public questions, please email genome@soe.ucsc.edu. All messages sent to that address are archived on a publicly accessible forum. If your question includes sensitive data, you may send it instead to genome-www@soe.ucsc.edu.